Be yourself; Everyone else is already taken.
— Oscar Wilde.
This is the first post on my new blog. I’m just getting this new blog going, so stay tuned for more. Subscribe below to get notified when I post new updates.
"Research is creating new knowledge." -Neil Armstrong
Be yourself; Everyone else is already taken.
— Oscar Wilde.
This is the first post on my new blog. I’m just getting this new blog going, so stay tuned for more. Subscribe below to get notified when I post new updates.
Finally finishing up the project. The extra week was sure needed to iron out all the kinks and make sure the documentation was completed properly. Also needed the time to make the presentation sucks we couldn’t do an actual presentation.
But even though this is the end of the project course this is only the start. I have learned so much from this project. Some of the things I learned late on can be applied to this very project to improve it. Would have been great to implement it all now but alas there are time limitation and prepping the data takes way too long to be experimental when we had a timeline.
We plan to continue working on the project once the semester is done to get the time series biased sorted and also get a Neural ODE model running. Can’t wait to return to this project later in the summer when I no longer would have other courses and no time restrictions. I can relax and enjoy the research like I hoped I would have during the project course.
Look forward for the next update! But until then stay safe and stay indoors!
After the long and tiring journey, our documentation and presentations are finally completed, the different comparisons on the final results using different datasets, gave us even more insights to the data and some ideas on what we can do to improve or to even test some hypothesis on sepsis prediction, but that’s a story for another day.
It was a great journey and for the moment it has come to an end. When our sanity returns we shall return to the project to improve and test some theories!
~~~Thank You for following our story~~~
We also made some modifications to the model by having the data undersampled and oversampled (with imbalance learn).
The results are shown here:
-> Random Undersampling
| Classifiers | F1 Score | Accuracy Score | Precision Score | Recall Score | ROC AUC Score |
| Random Guessing | 0.501245 | 0.501367 | 0.501347 | 0.501149 | 0.501358 |
| Logistic Regression | 0.944553 | 0.945775 | 0.966213 | 0.923844 | 0.945772 |
| Neural Network | 0.957754 | 0.957991 | 0.963063 | 0.952695 | 0.958023 |
| Random Forest | 0.995382 | 0.995362 | 0.99106 | 0.999742 | 0.995364 |
| Gaussian Naive Bayes | 0.87051 | 0.877277 | 0.920874 | 0.825597 | 0.877288 |
-> Oversampling with Imbalance Learn
| Classifiers | F1 Score | Accuracy Score | Precision Score | Recall Score | ROC AUC Score |
| Random Guessing | 0.498242 | 0.500363 | 0.497933 | 0.498552 | 0.500346 |
| Logistic Regression | 0.907205 | 0.910514 | 0.93717 | 0.879102 | 0.910364 |
| Neural Network | 0.928982 | 0.929131 | 0.926331 | 0.932271 | 0.929096 |
| Random Forest | 0.994949 | 0.994948 | 0.990014 | 0.999934 | 0.994972 |
| Gaussian Naive Bayes | 0.800169 | 0.820146 | 0.893598 | 0.72521 | 0.819687 |

Thank you for reading our journey from start to end of this project.
The new model works! It is great it still needs some tweaking but it is producing promising results. The scores are also better than the competition. We decided to compare it against different classifiers especially the ones the competition used in their research. Hopefully when they are tested they also perform well.
We have also been doing some of the documentation. That has been going smoothly. From the looks of it the data cleaning process we utilized seems to have made it easier for the models to classify if a patient has sepsis or not. However, we still need to test it by balancing the data since there are still more patients without sepsis that those with. Almost double actually.

We have decided to randomly under-sample the data and also look into methods from the imbalanced learn library to potentially over sample patients with sepsis.
The Feed Forward neural network performed great, with some tweaking and different approaches and data balancing, we had satisfactory results from the models. With results in hand the final grind is on to complete the documentation and presentations for the project now.
Hooray!! We finally got reasonable results from evaluating the PyTorch Feedforward Neural Network.
| Classifiers | F1 Score | Accuracy Score | Precision Score | Recall Score | ROC AUC Score |
| Random Guessing | 0.437825 | 0.500549 | 0.389329 | 0.500129 | 0.500473 |
| Logistic Regression | 0.931898 | 0.948762 | 0.964399 | 0.901518 | 0.940170 |
| Neural Network | 0.941330 | 0.955052 | 0.955112 | 0.928378 | 0.950201 |
| Random Forest | 0.995107 | 0.996177 | 0.990894 | 0.999355 | 0.996755 |
| Gaussian Naive Bayes | 0.862626 | 0.893530 | 0.865489 | 0.860305 | 0.887457 |
This is really good for us.

We finished up by adjusting hyperparameters.

Documentation and the final presentation were also touched up and results were added in.
We have decided to course correct with the guidance of our supervisor since we wouldn’t have the time to properly adjust an LSTM model with the time we have left. Especially checking we need to do a full write and and video presentation for the project.
We have decided to put the LSTM on hold as future work along with the Neural ODE. Definitely returning to conquer that hurdle once the semester is done.
We are instead using a feedforward model. This would require is to compress our time series data into single rows for each patient. Which wasn’t that hard to do we have started to write the code and that is going smoothly.
For each feature we will expand them to the following:
Hopefully this model will perform well and produce better results than our competition when trying to classifying sepsis.
We have also started to draft some of the write up so that things aren’t too rushed later on. Since it clear this is gonna be a close one.
Due to the nearing deadline an the absurd lengths and amounts of assignments and projects that courses have given us in place of our coursework evaluations time is running out and quickly.
Lacking a microwave from Steins Gate, a decision was made to delay the LSTM model as future works for the project, and continue using a Feed Forward Neural Network instead.
After building the LSTM in PyTorch, we continued to acquire terrible accuracy and F1 scores. This made us rethink our method of predicting sepsis.
We contacted our supervisor and decided to switch from an LSTM approach to using a Feedforward Neural Network instead.
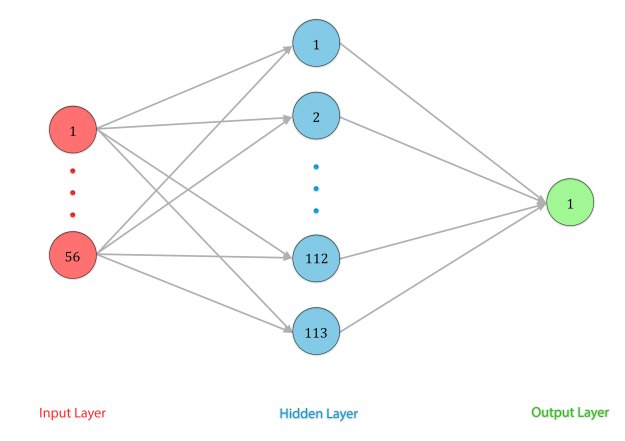
This would mean that we needed to re-create features.
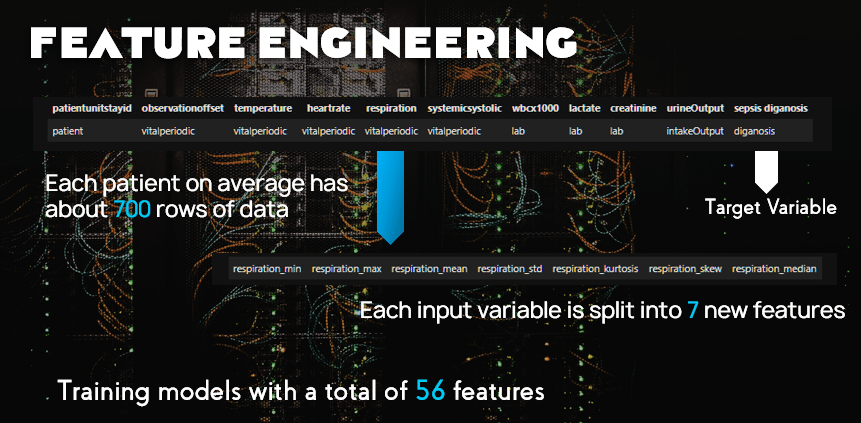

We took the time series data and made the multiple rows of data per patient into one row per patient.

We processed all of the full data into proper time series on the machine we got access it was so great seeing the code run. Took a good while though. The interpolate code took nine hours! I thought it would never end. But there is bad news….
The model didn’t produce good results. The f1 score is 0 which probably means the model is overfitting to the large amount of diagnosis of not sepsis since the accuracy score is close to 100%. It’s a bit depressing to be honest.

We also developed a model in pytorch that produced similar results and it take very long for the model to run so it’s not easy to tweak the model. Especially when there isn’t so much time left.
I also have a ton of other assignments to replace my course work exams since COVID 19 had to go ruin my final semester. This is overall further taking away from time I thought I would be able to spend on the project.
Hopefully things get better.